Circadian-Analysis
Cosinor Analysis and Circularized Kernel Density Estimation
C. William Yao
Source:vignettes/Circadian-Analysis.Rmd
Circadian-Analysis.RmdWhat will be covered in this tutorial? This specific tutorial is designed to show users how to use ActiGlobe to perform circadian analysis via cosinor models or Gaussian kernel density estimation (KDE). For data pre-processing or generation of graphic/excel report of daily actigraphy measures, please go to tutorials titled: Time-Shift and Graphic-Report.
Cosinor Analysis
After we segmented the data by date, we can now model the circadian
rhythm using the activity scores. For this, we will use the pre-travel
recording on the 27th of October. Since the recording was not
affected by the time change. We can simply store the unaffected segments
in a new variable called df, which stands for data
frame.
# Import data
data ("FlyEast")
BdfList <-
BriefSum (
df = FlyEast,
SR = 1 / 60,
Start = "2017-10-24 13:45:00",
TZ = "America/New_York"
)
# Let's extract actigraphy data from a single day
df <- BdfList$df
df <- subset (df, df$Date == "2017-10-27")Single-phase Model (i.e., single phase circadian rhythm)
Traditional Cosinor Model fitted via Ordinary Least Sqaure (OLS)
fit.ols <-
CosinorM (
time = df$Time,
activity = df$Activity,
tau = 24,
method = "OLS"
)
### Look at the parameters
fit.ols$coef.cosinor
#> MESOR Amplitude.24 Acrophase.24 Beta.24 Gamma.24
#> 177.297222 161.296243 -2.896002 -156.456357 -39.215893
### Plot
ggCosinorM (fit.ols)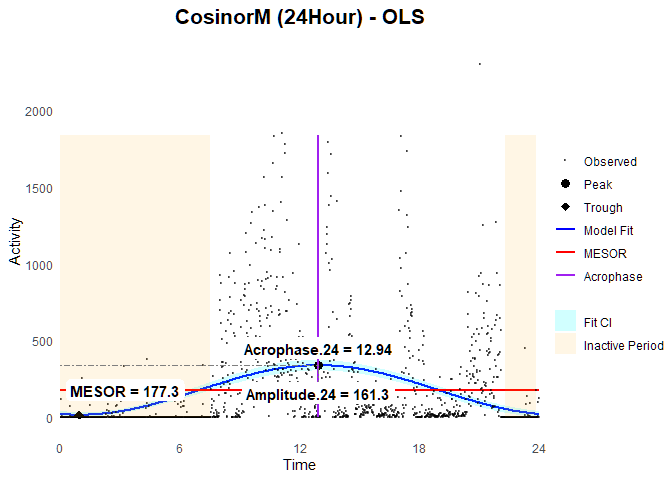
Note
If we process FlyEast using the
cosinor::cosinor.lm, we will notice that the results look a
bit different. This is because ActiGlobe treats the first time point of
the day as 00:00:00, whereas this is sometimes treated as
00:00:00 plus one unit of the device sampling rate due to
the use of the sequence generator.
Econometry-modified Cosinor Model with Feasible General Least Square (FGLS)
Besides, the traditional OLS-based cosinor
model,CosinorM also allows users to estimate the model
using the FGLS-based method. In ActiGlobe, the FGLS was implemented via
a weighted residual least square approximation. This two-step process
often produces a better representation of the “border” between sleep and
wakefulness than traditional OLS. Nevertheless, neither method can
realistically capture our activity patterns, given that they are simply
single-phase cosinor models.
fit.fgls <-
CosinorM (
time = df$Time,
activity = df$Activity,
tau = 24,
method = "FGLS"
)
### Look at the parameters
fit.fgls$coef.cosinor
#> MESOR Amplitude.24 Acrophase.24 Beta.24 Gamma.24
#> 181.169023 183.040124 -2.409542 -136.146165 -122.343405
### Plot
ggCosinorM (fit.fgls)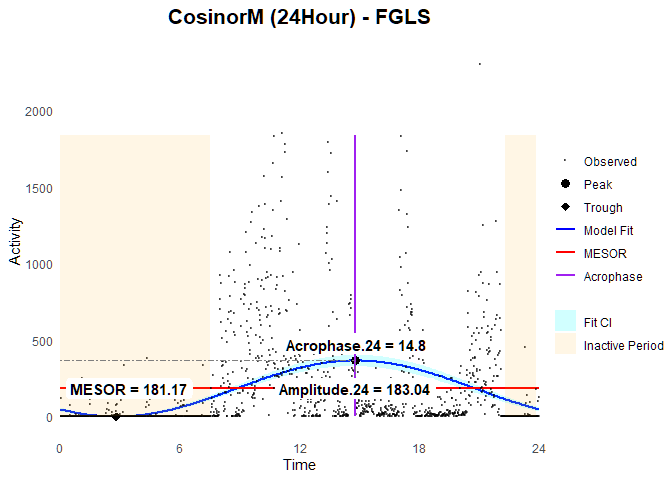
Dual-phase Model (i.e., 12 + 24hour rhythms)
While certain circadian measures, such as melatonin, often follow a
roughly 24-hour single-phase, human activity pattern is generally
fragmented. To capture the non-linearity in our activity pattern, it is
possible to add more phase components when using CosinorM.
Here, we provided two examples of cosinor models with two phase
components, each fitted using OLS or FGLS-based methods.
Traditional Cosinor Model fitted via Ordinary Least Sqaure (OLS)
fit.ols2 <-
CosinorM (
time = df$Time,
activity = df$Activity,
tau = c (12, 24),
method = "OLS"
)
fit.ols2$coef.cosinor
#> MESOR Amplitude.12 Amplitude.24 Acrophase.12 Acrophase.24 Beta.12
#> 177.2972222 136.4395529 161.2962432 -0.9644981 -2.8960017 77.7472728
#> Beta.24 Gamma.12 Gamma.24
#> -156.4563575 -112.1209756 -39.2158930
fit.ols2$post.hoc
#> MESOR.ph Bathyphase.ph.time Trough.ph Acrophase.ph.time
#> 182.629625 3.466667 -77.533671 10.783333
#> Peak.ph Amplitude.ph
#> 442.792922 260.163297
### Plot
ggCosinorM (fit.ols2)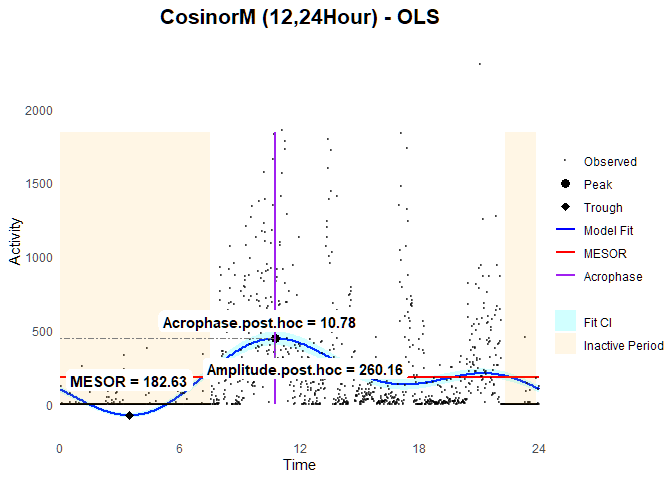
Eocnometry-modified Cosinor Model with Feasible General Least Square (FGLS)
fit.fgls2 <-
CosinorM (
time = df$Time,
activity = df$Activity,
tau = c (12, 24),
method = "FGLS"
)
fit.fgls2$coef.cosinor
#> MESOR Amplitude.12 Amplitude.24 Acrophase.12 Acrophase.24 Beta.12
#> 148.6450319 52.7954160 128.4419382 -0.7394464 -2.7311777 39.0074572
#> Beta.24 Gamma.12 Gamma.24
#> -117.7755191 -35.5777211 -51.2470351
fit.fgls2$post.hoc
#> MESOR.ph Bathyphase.ph.time Trough.ph Acrophase.ph.time
#> 148.959604 3.450000 -8.218935 11.750000
#> Peak.ph Amplitude.ph
#> 306.138144 157.178540
### Plot
ggCosinorM (fit.fgls2)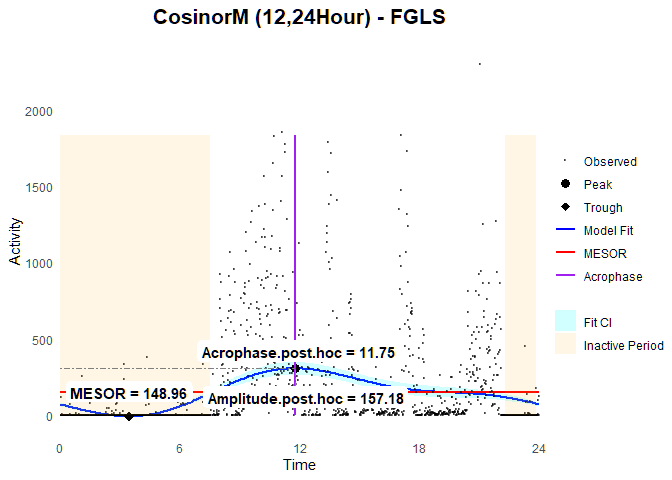
It is worth noting that we cannot individually interpret each cosinor parameter alone when fitting a multicomponent model. This is because the rest-activity patterns estimated by general linear regression are the cumulative sums of all the fitted cosinor components (i.e., both 12- and 24-hour waves). As such, it is better to use the post hoc-extracted parameters for analysis.
#> Beta.12 Gamma.12 phi.12
#> 77.74727 -112.12098 10.15794
#> Beta.24 Gamma.24 phi.24
#> -156.45636 -39.21589 12.93809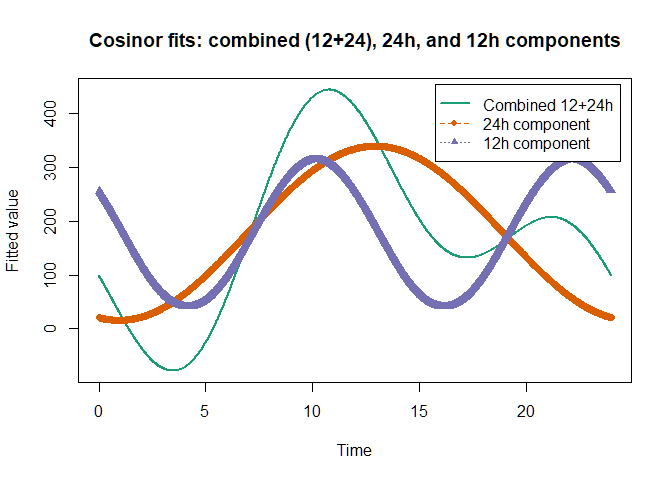
Gaussian Kernel Density Estimation on Circularized Time
Besides the traditional cosinor rhythmetry, ActiGlobe also offers
Gaussian kernel density estimation (KDE). In contrast to the traditional
cosinor method, CosinrM.KDE uses nonparametric kernel
smoothing to map the activity pattern on a circularized time domain.
This strategy allows it to better respect the intensity and the duration
of the underlying rest-activity patterns, without forcing a sinusoidal
shape. To ease transition from cosinor model, we designed
CosinorM.KDE to generate cosinor-like parameters by
projecting the model onto an imaginary cosinor plane.
fit.KDE <-
CosinorM.KDE (
time = df$Time,
activity = df$Activity
)
### Look at the parameters
fit.KDE$coef.cosinor
#> MESOR Amplitude.24 Acrophase.24 Beta.24 Gamma.24
#> 177.327801 157.744980 -2.895821 -153.004740 -38.380048
fit.KDE$post.hoc
#> MESOR.ph Bathyphase.ph.time Trough.ph Acrophase.ph.time
#> 296.160217 5.433333 9.099325 10.483333
#> Peak.ph Amplitude.ph
#> 583.221110 287.060893
ggCosinorM (fit.KDE)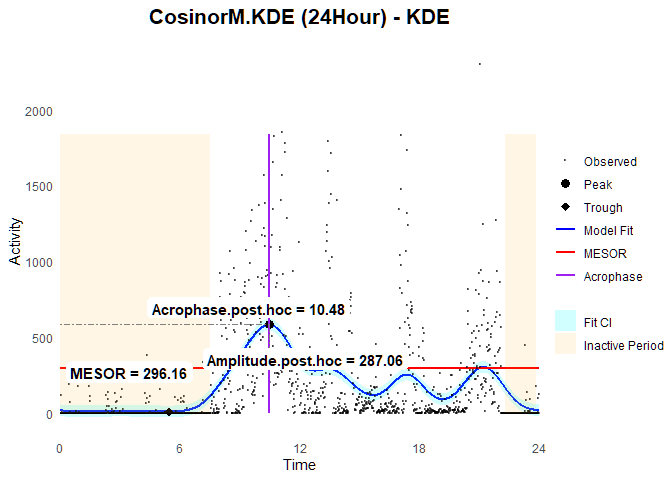
It is worth noting that while CosinorM.KDE does provide
a variance and standard error for the fitted rest-activity pattern; this
is a density‑based signal‑processing strategy, not a regression model.
As a signal‑processing technique, CosinorM.KDE lacks the
ability to directly provide standard errors or confidence intervals for
cosinor parameters and related post hoc estimates (e.g., bathyphase).
Thankfully, we can use bootstrap resampling to empirically estimate
confidence intervals and standard errors for these post‑hoc parameters.
Note that CosinorM.KDE is designed to stabilize estimates
when data are irregularly spaced or limited to short recording
intervals.
boot.seci (
object = fit.KDE,
ci_level = 0.95,
n = 100
) ### for demonstration, the number of bootstraps was limited to 100.
#> Estimate Std Error t value 2.5% 97.5%
#> MESOR 177.30 4.60 38.56 168.99 185.45
#> Amplitude.24 157.54 7.15 22.06 145.39 171.51
#> Acrophase.24 -2.89 0.04 -73.15 -2.96 -2.82
#> Beta.24 -152.35 7.90 -19.38 -167.59 -138.84
#> Gamma.24 -39.61 5.28 -7.27 -48.31 -29.42
#> MESOR.ph 295.55 14.50 20.43 266.18 324.46
#> Bathyphase.ph.time 4.35 1.83 2.96 0.60 6.13
#> Trough.ph 8.12 1.84 4.93 5.09 11.66
#> Acrophase.ph.time 10.50 0.31 33.60 9.97 11.05
#> Peak.ph 582.97 29.10 20.04 522.64 638.85
#> Amplitude.ph 287.43 14.66 19.58 257.36 314.75Currently, was designed to compute standard errors and confidence intervals in serial processes, which means that it will take some time to finish the computation.
Visualized Model Comparisons
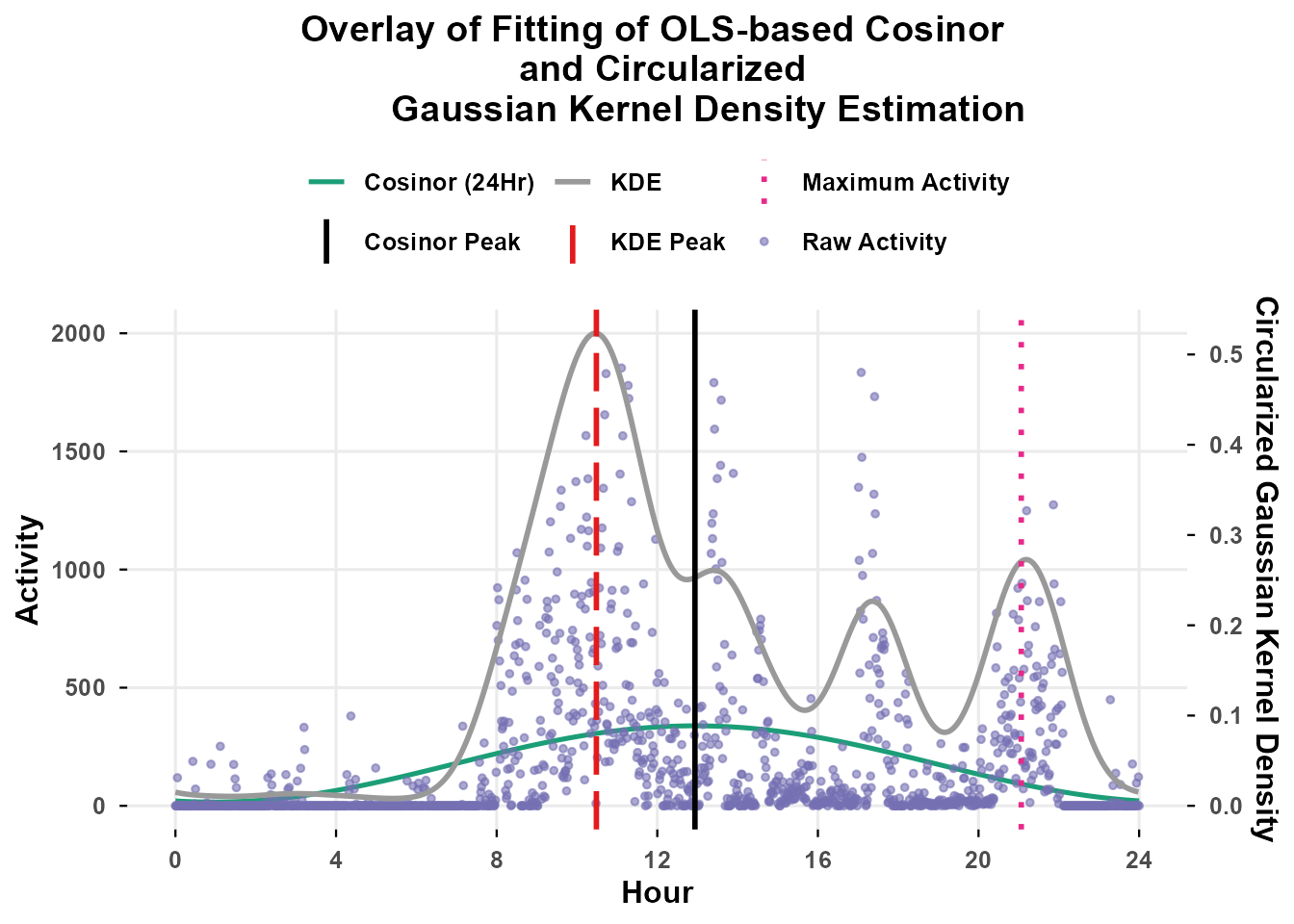
OLS-Cosinor vs. Kernel Density Estimation
When we compare the KDE-based estimate with the OLS-based cosinor
model, we can immediately see that CosinorM.KDE better
captures the complexity of the wearer’s activity pattern. Since
CosinorM.KDE does not assume a specific pattern of
activity, it avoids introducing bias from assumptions or model
specifications into the rest-activity-based circadian parameters.